Classified by application
All Products
Signaling Pathways
- Angiogenesis/Protein Tyrosine Kinase
- Apoptosis Pathway
- Cell Cycle/DNA Damage
- Epigenetics
-
GPCR/G Protein
- Urotensin Receptor
- Melatonin Receptor
- Neurotensin Receptor
- Glucose Transporter
- Motilin Receptor
- TGR5
- Cholecystokinin Receptor
- Adenylate Cyclase
- Somatostatin Receptor
- Oxytocin Receptor
- Bradykinin Receptor
- Guanylate Cyclase
- Melanocortin Receptor
- Imidazoline Receptor
- Neuropeptide Receptor
- PAR
- CRFR
- Sigma Receptor
- S1P Receptor
- GHSR
- PKA
- OX Receptor
- Adenosine Receptor
- Glucagon Receptor
- Prostaglandin Receptor
- NK Receptor
- GnRH Receptor
- Leukotriene Receptor
- Glucocorticoid Receptor
- mAChR
- P2 Receptor
- GPR
- CGRP Receptor
- CXCR
- Histamine Receptor
- CaSR
- Ras/Rho
- Vasopressin Receptor
- Smoothened/Smo
- mGluR
- Opioid Receptor
- Dopamine Receptor
- Cannabinoid Receptor
- Angiotensin Receptor
- Adrenergic Receptor
- SGLT
- Endothelin Receptor
- CCR
- 5 HT Receptor/Serotonin Receptor
- Hormone Pathay
- Ion Channel/Membrane Transporter
- Jak/Stat Pathway
- MAPK Pathway
- Microbiology/Virology
-
Neuro Signaling Pathway
- Imidazoline Receptor
- Monoamine Oxidase
- COMT
- Amyloid β
- NK Receptor
- Beta secretase
- AChE
- SSRI
- CaMK
- P glycoprotein
- Opioid Receptor
- Dopamine Receptor
- Adrenergic Receptor
- Histamine Receptor
- mGluR
- Gamma secretase
- Dopamine Transporter
- 5 HT Receptor/Serotonin Receptor
- P2 Receptor
- FAAH
- CGRP Receptor
- nAChR
- COX
- GABA Receptor
- mAChR
- NFκB
- PI3K/Akt/mTOR
-
Protease/Metabolic Enzyme
- COMT
- 11β HSD
- Carboxypeptidase
- ACC
- Xanthine Oxidase
- Pyruvate Kinase
- Mitochondrial Metabolism
- ALDH
- SCD
- Serine Protease
- MLR
- FTase
- CETP
- Neprilysin
- MAGL
- Carbonic Anhydrase
- E1 E2 E3 Enzyme
- Elastase
- PAI1
- NOX
- ROR
- Glucokinase
- Aldose Reductase
- Tyrosinase
- FAS
- DGAT
- Lipoxygenase
- Nampt
- IDO
- IDH
- MMP
- Cathepsin
- Phosphatase
- Thrombin
- FXR
- RAR/RXR
- HMGCR
- FAAH
- Gamma secretase
- RAAS/ACE
- Caspase
- HIV Protease
- Aminopeptidase
- HCV Protease
- HSP
- Tryptophan Hydroxylase/TPH
- Factor Xa
- DPP
- Procollagen C Proteinase
- Integrase
- Phosphodiesterase/PDE
- Phospholipase
- LXR
- Proteasome
- Cytochrome P450
- TGF beta/Smad
- Wnt/Stem Cell
- Others Pathway
Research Areas
- Metabolic Disease
- Neurological Disease
- Endocrinology
- Diabetes
- Cardiovascular Disease
- Immunology
- Allergy
- Inflammation
- Infection
- Cancer
Nature products
Antibodies
Peptides
Catalysts
Impurities
Intermediate
- Amino acid
- Ammonia Series
- Azaindazole
- Pyridazine Series
- Nucleoside Series
- Quinazoline
- Pyrazole Series
- Boric acid
- Pyrimidine Series
- Quinoline Series
- Pyrrole Series
- Thiophene Series
- Saccharide
- Azaindole
- Indole Series
- Piperidine Series
- Indazole Series
- Heterocyclic Series
- Benzene Series
- Pyridine Series
Raw Materials
Epigenetic Reader Domain
| Chemical Structure | Cat. No. | Product Name | CAS No. |
|---|
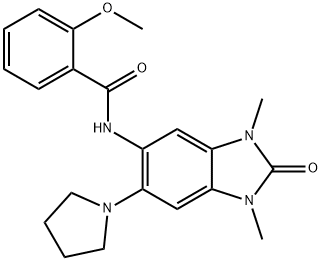 |
BCP23818 | PFI-4 New | 900305-37-5 |
PFI-4 is a potent and highly selective BRPF1 Bromodomain (BRPF1B) inhibitor, with an IC50 of 172 nM. PFI-4 can be used to explore the functional mechanisms of the HBO1/BRPF1 complex and to study bone loss and osteolytic malignant bone lesions.
|
|||
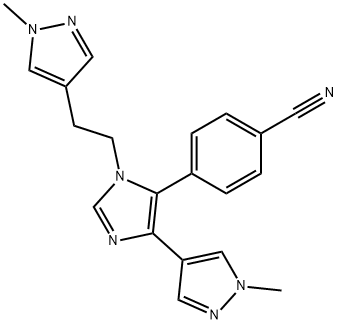 |
BCP23999 | BAZ2-ICR New | 1665195-94-7 |
BAZ2-ICR is a potent, selective, cell active and orally active BAZ2A/B bromodomains inhibitor with IC50s of 130 nM and 180 nM, and Kds of 109 nM and 170 nM, respectively. BAZ2-ICR shows 10-15-fold selectivity for binding BAZ2A/B over CECR2 and >100-fold selectivity over all other bromodomains. BAZ2-ICR is an epigenetic chemical probe.
|
|||
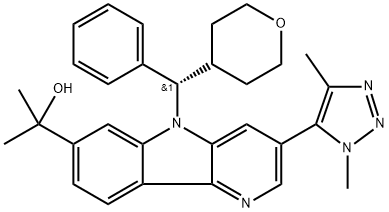 |
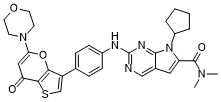
BCP25147 | BMS 986158 New | 1800340-40-2 |
BMS-986158 is an inhibitor of the bromodomain and extra-terminal (BET) proteins.
|
|||
 |
BCP48567 | SRX3177 New | 2241237-51-2 |
SRX3177 is a novel potent triple action inhibitor, targeting BRD, PI3K, and CDK.
|
|||
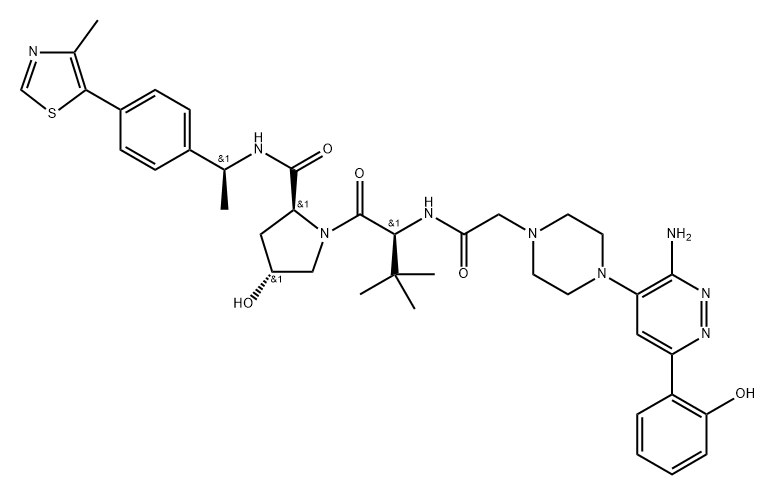 |
BCP46987 | AU-15330 New | 2380274-50-8 |
AU-15330 is a novel, highly specific and VHL-dependent PROTAC degrader of SWI/SNF ATPase components (SMARCA2, SMARCA4 and PBRM1) that shows preferential cytotoxicity in enhancer-binding transcription factor-addicted cancers at low nanomolar concentrations.
|
|||
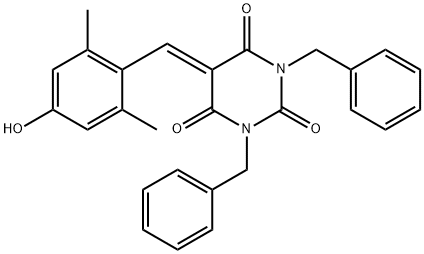 |
BCP46797 | EML 425 New | 1675821-32-5 |
EML425 is a potent and selective CREB binding protein (CBP)/p300 inhibitor with IC50s of 2.9 and 1.1 μM, respectively.
|
|||
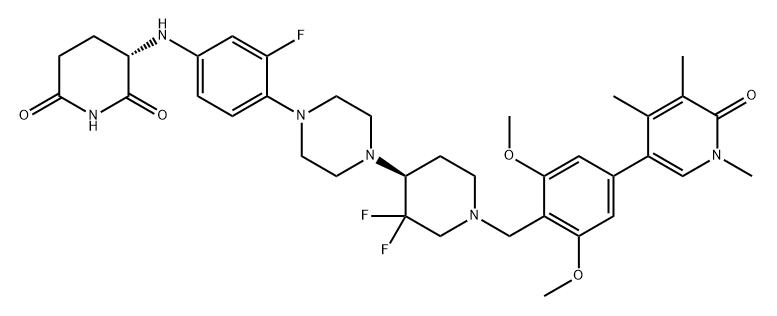 |
BCP46564 | CFT-8634 New | 2704617-96-7 |
CFT-8634 is a degrader targeting BRD9 extracted from patent WO2021178920A1 compound 173.
|
|||
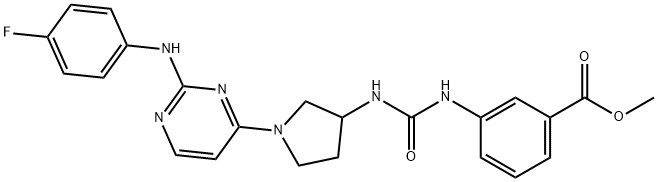 |
BCP46513 | GSK1379725A New | 1802251-00-8 |
GSK1379725A is the first selective inhibitor of BPTF over Brd4.
|
|||
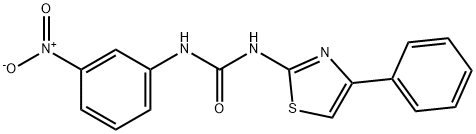 |
BCP46449 | BAZ1A-IN-1 New | 941521-45-5 |
BAZ1A-IN-1 is a potent inhibitor of BAZ1A (bromodomain-containing protein). BAZ1A-IN-1 shows a KD value of 0.52 μM against BAZ1A bromodomain.
|
|||
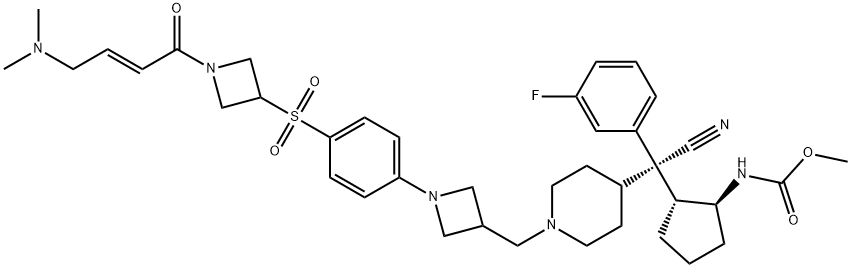 |
BCP46311 | M-525 New | 2173582-08-4 |
M-525 is a first-in-class, highly potent, irreversible small-molecule inhibitor of the menin-MLL interaction.
|
|||